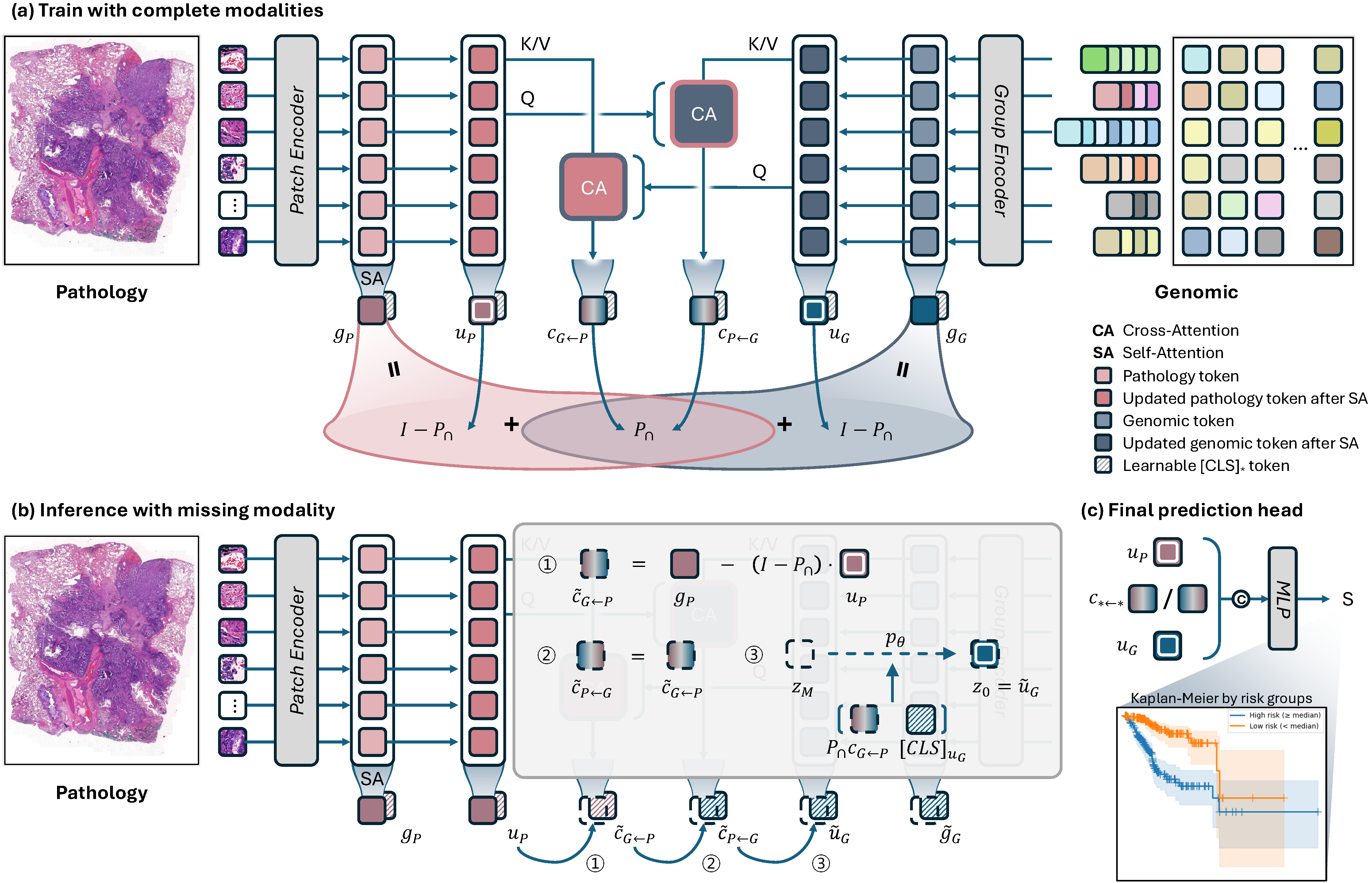
Overall architecture of MUST. The framework extracts global representations via self-attention, computes shared information through bidirectional cross-attention, and decomposes each modality into modality-specific and shared components through algebraic constraints in a learned low-rank subspace. Missing modality-specific components are reconstructed via conditional latent diffusion models.